- Title
-
Fine-tuning amyloid precursor protein expression through non-sense mediated mRNA decay
- Authors
- Rahmati, M., Chebli, J., Banote, R.K., Roselli, S., Agholme, L., Zetterberg, H., Abramsson, A.
- Source
- Full text @ eNeuro
|
|
|
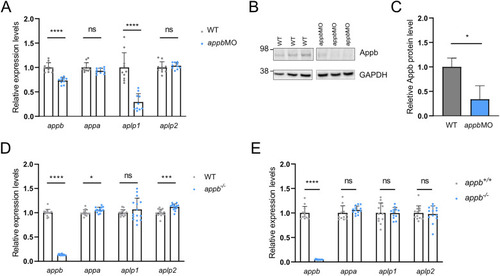
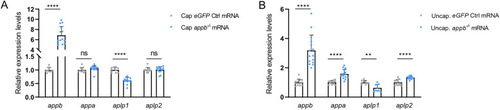
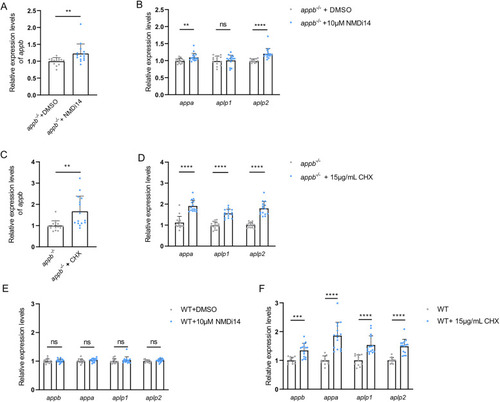
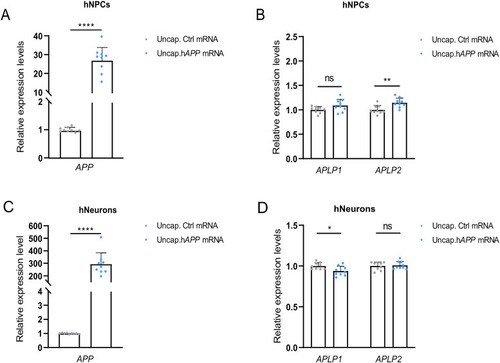
Relative gene expression in translation-blocking |
|
Expression of |
|
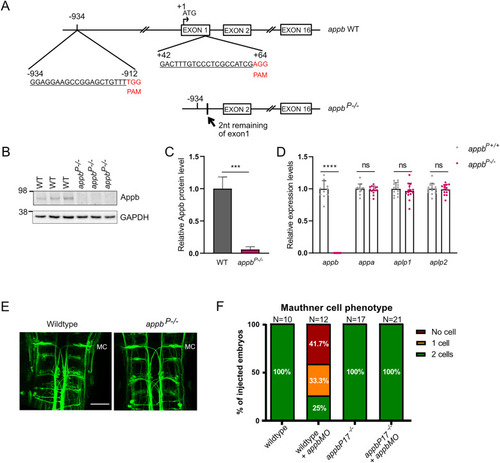
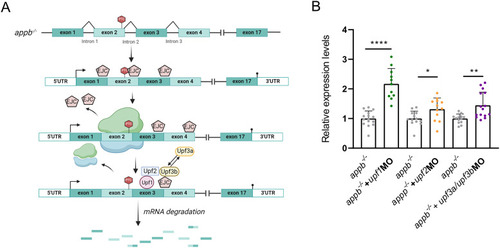
The RNA-less |
|
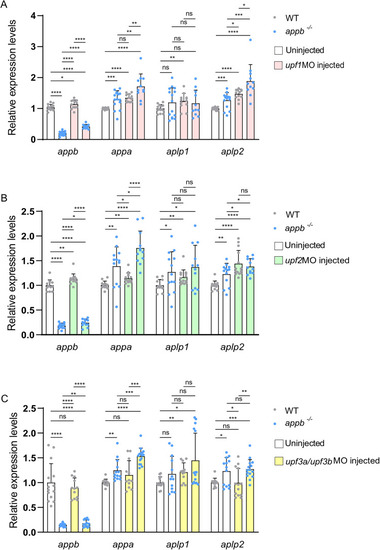
Expression of |
|
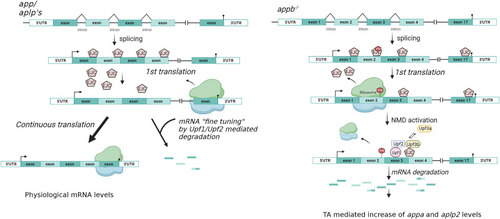
NMD pathway in |
|
Expression of |
|
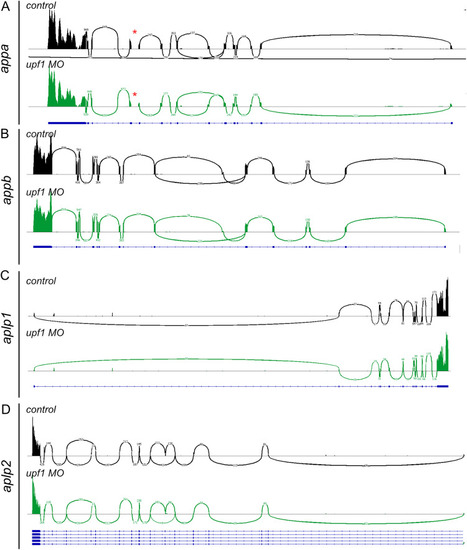
Sashimi plots of |
|
TA in hNPCs but not in terminally differentiated neuron cells transfected with unstable h |
|
NMD pathway. |