Figure 1
- ID
- ZDB-FIG-230428-24
- Publication
- Yamaguchi et al., 2023 - Calaxin stabilizes the docking of outer arm dyneins onto ciliary doublet microtubule in vertebrates
- Other Figures
- All Figure Page
- Back to All Figure Page
|
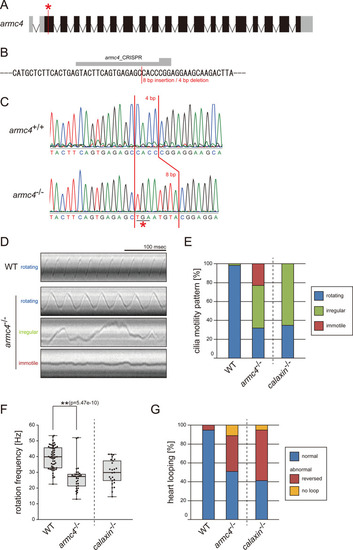
(A) Genomic organization of zebrafish armc4 gene. Black boxes: exons. Gray boxes: untranslated regions. Red asterisk: the genome-editing target site. (B) CRISPR/Cas9 target sequence. (C) Sanger sequencing of armc4+/+ and armc4−/− fish around the genome-editing target site. The 8 bp-insertion in armc4−/− includes a stop codon (red asterisk). (D) Typical kymographs of Kupffer’s vesicle cilia in WT and armc4−/− embryos. Kymograph patterns were categorized into three classes: rotating (blue), irregular (green), and immotile (red). Scale bar: 100 ms. (E) Ratios of each motility class. Number of cilia: 58 (WT) and 100 (armc4−/−). (F) Rotational frequencies of Kupffer’s vesicle cilia. Number of cilia: 58 (WT) and 32 (armc4−/−). Boxes correspond to the first and third quartiles, lines inside the boxes indicate the medians, and whiskers extend to the full range of the data. p-Value was calculated with Welch’s t-test. (G) Directions of heart looping. Number of embryos: 110 (WT) and 63 (armc4−/−). For comparison, calaxin-/- data from Sasaki et al., 2019 are displayed in (E, F, and G).
|