- Title
-
Environmental chemicals affect circadian rhythms: An underexplored effect influencing health and fitness in animals and humans
- Authors
- Zheng, X., Zhang, K., Zhao, Y., Fent, K.
- Source
- Full text @ Environ. Int.
|
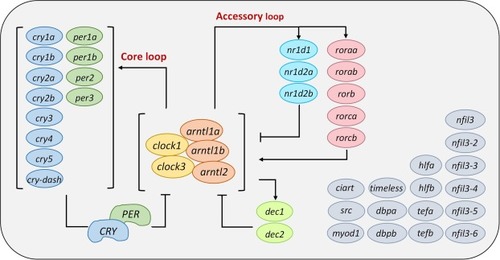
Circadian rhythm system in zebrafish and associated genes in the core and accessory loops. Our homologous alignments (based on current zebrafish genome annotation (GRCz11) in Ensembl database) identified 43 circadian genes. Multiple paralogues exist in zebrafish owing to the evolutionary whole-genome duplication events. Full name of each gene is shown in the Supplementary Material in Table S1. The core component is the clock/arntl heterodimer, which binds in the promoters of per and cry genes to induce its transcription, and the physical binding of PER and CRY proteins will interact to inhibit clock/arntl heterodimer transcription, thereby forming a core negative feedback loop. Nuclear receptor genes, nr1ds and rors, are also involved in as accessory loops. The former is regulated by a negative and the latter by a positive feedback regulation mechanism. Circadian genes in grey color indicate that their mechanisms of action are not yet entirely understood. |
|
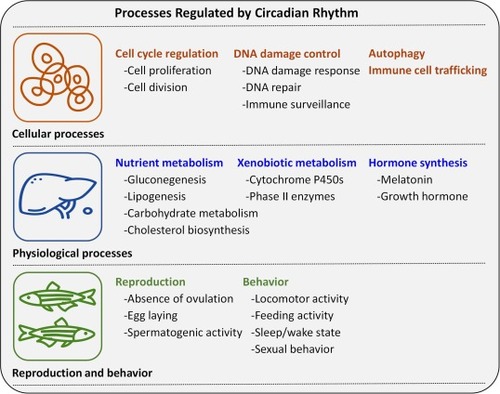
Physiological processes at different levels of biological organization that are regulated by circadian rhythms. They can be affected by circadian disrupters. |
|
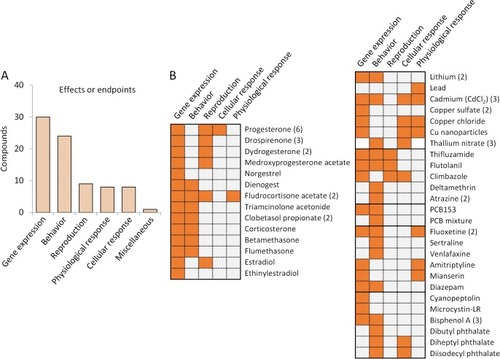
Summary of circadian related effects assessed in environmental chemicals. (A) Number of chemicals with circadian gene changes at different effect levels. Categories are gene expression or cellular, physiological, reproductive and behavioral effects. (B) Details of reported effects on the endpoints for each circadian disrupter, grouped in steroid hormones and other compounds. The summary of effects for each compound derived from different studies (number of studies given in parenthesis) is shown. More information is shown in Table 1 and Table S2. |