- Title
-
Structure/Function Studies of the α4 Subunit Reveal Evolutionary Loss of a GlyR Subtype Involved in Startle and Escape Responses
- Authors
- Leacock, S., Syed, P., James, V.M., Bode, A., Kawakami, K., Keramidas, A., Suster, M., Lynch, J.W., Harvey, R.J.
- Source
- Full text @ Front. Mol. Neurosci.
|
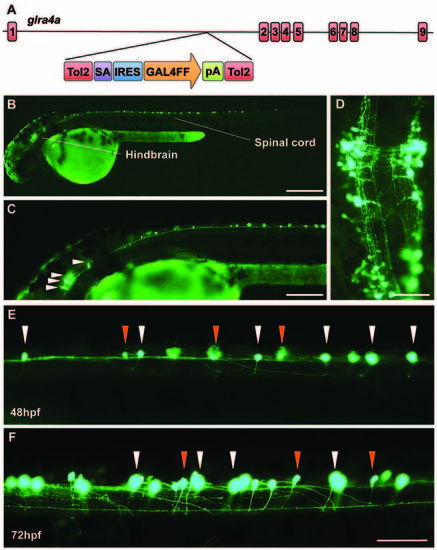
A novel zebrafish glra4a gene trap reveals expression in the zebrafish brain and spinal cord. (A) The zebrafish SAIGFF16B line contains the gene trap construct T2KSAGFF inserted between exons 1 and 2 of glra4a. This cassette contains a splice acceptor (SA), followed by an internal ribosome entry site (IRES), the coding region for the Gal4FF transcription activator and a polyadenylation site (pA). Gal4FF expression in SAIGFF16B was visualized by creating double transgenic fish carrying the Gal4FF transgene and a GFP reporter gene placed downstream of the Gal4-recognition sequence (UAS:GFP). (B,C) GFP expression in the glra4a gene-trap line at 48 hours post fertilization (hpf) reveals that glra4a is predominantly expressed four clusters of hindbrain commissural neurons and selected spinal commissural interneurons. (D) High magnification dorsal view of neurons indicated by arrows in (C). (E,F) Images show a portion of the zebrafish spinal cord at 48 hpf (E) and 72 hpf (F) showing glra4a expression in commissural primary (CoPA, white arrows) and secondary (CoSA, orange arrows). Scale bars: (B): 500 μm; (C); 250 μm; (D): 150 μm; (E,F): 50 μm. EXPRESSION / LABELING:
|
|
Morpholino oligonucleotides (SMO1 and SMO2) block splicing of glra4a. (A) Gross morphology of wild-type embryos and those injected with GlyR α4a translation-blocking (TMO) or splice site (SMO1 and SMO2) morpholinos. (B) Schematic of RT-PCR analysis of glra4a morpholinos confirmed the deletion of exon 7 for α4a-SMO1 (lane 3) and the deletion of part of exon 7 for α4a-SMO2 (lane 4). Intact GlyR α4a exon 7–9 PCR products are seen in the control (393 bp, lane 2) and smaller amounts are also observed in lanes 3 and 4. A fragment of around 178 bp is seen in SMO1, which is made up of exon 6 and 8 only. A fragment of 321 bp in SMO2 contains exons 6 and 8 and part of exon 7, which excludes the region that codes for TM2. GlyR α1 exon 7 is present in control and both SMO fish (lower panel). (C,D) DNA sequencing of GlyR α4a cDNAs from SMO1 and SMO2 treated fish reveals that SMO1 results in a 215 bp deletion, resulting in a frameshift and premature stop codon before M1, while SMO2 results in a 72 bp deletion and loss of 24 amino acids, including the majority of the pore-forming M2 domain. PHENOTYPE:
|
|
Embryos injected with α4a-SMO1 and the α4aR278Q mutant embryos fail to exhibit correct escape behavior. (A) Wild-type (control) fish responds to touch by with a C-bend, a turn away from the stimulus and rapid swimming. However, embryos injected with α4a-SMO1 (B) or with mRNA for the GlyR α4aR278Q subunit (C) respond more slowly, and have a pronounced head retraction—similar to a localized beo-like contraction (arrow). Single images were extracted from high-speed movies of α4a-SMO1 morphants and embryos injected with GlyR α4aR278Q subunit mRNA. PHENOTYPE:
|