- Title
-
Maximizing CRISPR/Cas9 phenotype penetrance applying predictive modeling of editing outcomes in Xenopus and zebrafish embryos
- Authors
- Naert, T., Tulkens, D., Edwards, N.A., Carron, M., Shaidani, N.I., Wlizla, M., Boel, A., Demuynck, S., Horb, M.E., Coucke, P., Willaert, A., Zorn, A.M., Vleminckx, K.
- Source
- Full text @ Sci. Rep.
|
Theoretical models of how gRNA-specific efficiencies and frameshift gene editing outcome probabilities influence the cellular composition and percentage of protein knockout cells in a mosaic F0 animal model. ( |
|
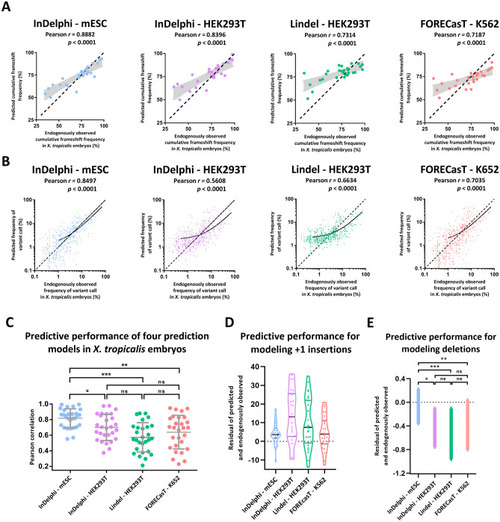
The InDelphi prediction model, trained in mESC cells, accurately predicts CRISPR/Cas9 gene editing outcomes and outperforms several other prediction models in |
|
The InDelphi-mESC model accurately predicts CRISPR/Cas9 gene editing outcomes in |
|
Integrating CRISPRscan and the InDelphi-mESC model allows identification of efficient high frameshift frequency gRNAs in |