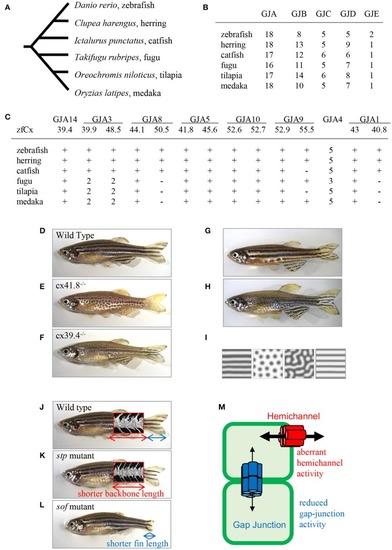
Connexins in teleosts. (A) Phylogenic relationship among six teleost species (Chen et al., 2004). (B) The number of connexin-encoding genes in six teleost species. (C) The number of genes encoding connexins of the alpha family gap junctions. “zfCx” indicates zebrafish connexins; “+” and “−” indicate the existence or absence, respectively, of an ortholog. If more than one orthologous gene was found, gene numbers are indicated (B,C; Connexin sequences were obtained from genome data base in Sanger Institute, http://www.sanger.ac.uk/). (D–I) Connexins in zebrafish pigment patterns. Wild-type zebrafish (D; Watanabe and Kondo, 2012); leopard mutant (E; Watanabe and Kondo, 2012); luchs mutant (F; Irion et al., 2014; Watanabe et al., 2016); transgenic zebrafish Tg(mitfa-cx41.8) >> leopard (G; Watanabe and Kondo, 2012); transgenic zebrafish Tg(mitfa-cx41.8M7) >> wild-type (H; Watanabe and Kondo, 2012); reaction-diffusion (R-D) patterns (I; Watanabe and Kondo, 2012). (J–M) Connexins in zebrafish bones; micro-CT images of vertebrae are superimposed. Wild-type zebrafish (J; Misu et al., 2016), stp mutant (K; Misu et al., 2016), sof mutant (L; Iovine et al., 2005; Misu et al., 2016). Schematic presentation of gap junction and hemichannel functions in zebrafish mutants (M; Misu et al., 2016). Red font, functional activity of hemichannels in the stp-Cx43 mutant; blue font, functional activity of gap junctions in the sof-Cx43 mutant.
|