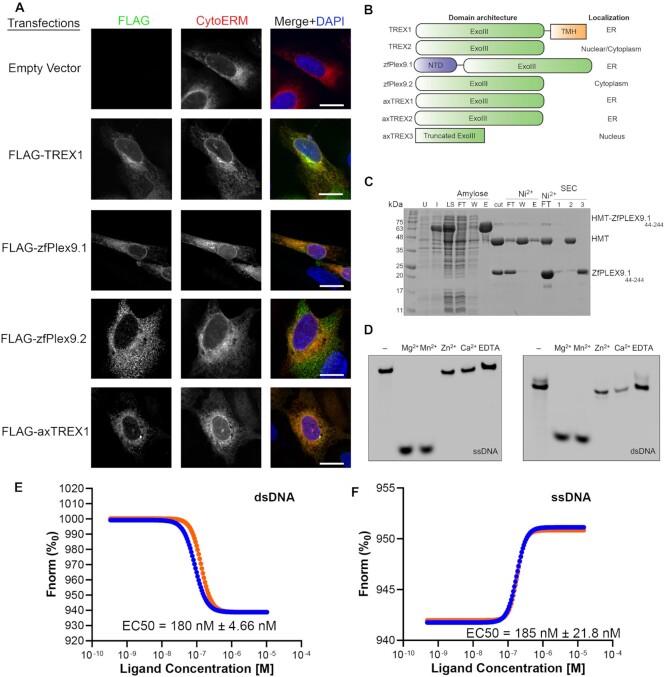
Fig. 4
Plex9 proteins represent a novel class of DEDDh DNA exonucleases. (A) Ectopic expression of zebrafish Plex9.1/.2 (zfPlex9.1/.2) and axTREX1/2 reveals that these proteins localize to the ER and cytoplasm. Proteins were FLAG-tagged and expressed in U2OS cells to assess their localization. A domain of Cytochrome p450 (CytoERM) tagged to mScarlet (addgene #85066) was utilized as an ER marker. An empty vector (BlueScript) was used as a control. Scale bars represent 10 μm. (B) The different domain architectures of TREX1/2 and Plex9 proteins and their known/determined localizations are shown on the right. TREX2 localization was assessed previously as both nuclear and cytoplasmic. (C) Purification of zfPlex9.1. zfPlex9.1 was expressed as a fusion to sequences encoding for hexahistidine, maltose binding protein, and a TEV protease cleavage site (HMT). SDS-PAGE analysis is shown for samples of uninduced (U) and induced (I) cells, soluble lysate (LS), after TEV cleavage (cut), flowthrough (FT), wash (W), and elution (E) fractions of amylose and Ni2+ affinity chromatography, and fractions from size exclusion chromatography (SEC). (D) zfPlex9.1 requires Mg2+ or Mn2+ for activity. Fluorescent oligonucleotides were incubated with zfPlex9 and the indicated metal. Samples were resolved by UREA-PAGE and visualized via fluorescence imaging. ‘-’ refers to input without the addition of cations. (E, F) DNA binding of zfPlex9.1. Microscale thermophoresis was used to quantify the affinity between zfPlex9.1 with (E) ssDNA and (F) dsDNA (n = 2). The different coloured lines are data shown for two replicates.